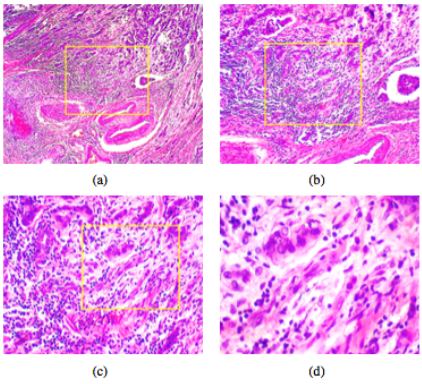
The Breast Cancer Histopathological Image Classification (BreakHis) is composed of 9,109 microscopic images of breast tumor tissue collected from 82 patients using different magnifying factors (40X, 100X, 200X, and 400X). To date, it contains 2,480 benign and 5,429 malignant samples (700X460 pixels, 3-channel RGB, 8-bit depth in each channel, PNG format). This database has been built in collaboration with the P&D Laboratory – Pathological Anatomy and Cytopathology, Parana, Brazil (http://www.prevencaoediagnose.com.br). We believe that researchers will find this database a useful tool since it makes future benchmarking and evaluation possible.
Characteristics
The dataset BreaKHis is divided into two main groups: benign tumors and malignant tumors. Histologically benign is a term referring to a lesion that does not match any criteria of malignancy – e.g., marked cellular atypia, mitosis, disruption of basement membranes, metastasize, etc. Normally, benign tumors are relatively “innocents”, presents slow growing and remains localized. Malignant tumor is a synonym for cancer: lesion can invade and destroy adjacent structures (locally invasive) and spread to distant sites (metastasize) to cause death.
In current version, samples present in dataset were collected by SOB method, also named partial mastectomy or excisional biopsy. This type of procedure, compared to any methods of needle biopsy, removes the larger size of tissue sample and is done in a hospital with general anesthetic.
The BreaKHis 1.0 is structured as follows:
| Magnification | Benign | Malignant | Total |
|---|---|---|---|
| 40X | 652 | 1,370 | 1,995 |
| 100X | 644 | 1,437 | 2,081 |
| 200X | 623 | 1,390 | 2,013 |
| 400X | 588 | 1,232 | 1,820 |
| Total of Images | 2,480 | 5,429 | 7,909 |
Both breast tumors benign and malignant can be sorted into different types based on the way the tumoral cells look under the microscope. Various types/subtypes of breast tumors can have different prognoses and treatment implications. The dataset currently contains four histological distinct types of benign breast tumors: adenosis (A), fibroadenoma (F), phyllodes tumor (PT), and tubular adenona (TA); and four malignant tumors (breast cancer): carcinoma (DC), lobular carcinoma (LC), mucinous carcinoma (MC) and papillary carcinoma (PC).
Each image filename stores information about the image itself: method of procedure biopsy, tumor class, tumor type, patient identification, and magnification factor. For example, SOB_B_TA-14-4659-40-001.png is the image 1, at magnification factor 40X, of a benign tumor of type tubular adenoma, original from the slide 14-4659, which was collected by procedure SOB. More formally, the format of image file name is given by the following BNF notation:
<BIOPSY_PROCEDURE>_<TUMOR_CLASS>_<TUMOR_TYPE>-<YEAR>-<SLIDE_ID>-<MAG>-<SEQ>
<BIOPSY_PROCEDURE>::=SOB
<TUMOR_CLASS>::=M|B
<TUMOR_TYPE>::=<BENIGN_TYPE>|<MALIGNANT_TYPE>
<BENIGN_TYPE>::=A|F|PT|TA
<MALIGNANT_TYPE>::=DC|LC|MC|PC
<YEAR>::=<DIGIT><DIGIT>
<PATIENT_ID>::=<NUMBER><SEC>
<SEQ>::=<NUMBER>
<MAG>::=40|100|200|400
<NUMBER>::=<NUMBER><DIGIT>|<DIGIT>
<SEC>::=<SEC>::<LETTER>|<LETTER>
<DIGIT>::=0|1|…|9
<LETTER>::=A|B|…|Z
How to obtain access to the images
The BreakHis Database may be used for non-commercial research provided you acknowledge the source of the image by citing the following paper in publications about your research:
You may download the BreaKHis database using this link:
http://www.inf.ufpr.br/vri/databases/BreaKHis_v1.tar.gz
We kindly ask you to fill the following form after downloading the dataset.
- decompress the file mkfold.tag.gz
- copy the file BreakHis_v1.tar.gz into the mkfold directory
- decompress the BreakHis_v1.tar.gz file
- run the script <python mkfold.py>
It will create five directories inside the mkfold directory containing the structure used in [1].
References:
- [2] Spanhol, F., Oliveira, L. S., Petitjean, C., and Heutte, L., Breast Cancer Histopathological Image Classification using Convolutional Neural Network, International Joint Conference on Neural Networks (IJCNN 2016), Vancouver, Canada, 2016.
- Caffe_models.tar.gz – It contains two Caffe model definition files (protobuf model format), a solver file and a trained Caffe model. This model is snapshot of iteration 80,000, considering the strategy #4, fold 1 magnification factor 40x, as described in [1]. Please, edit the files and adjust the paths properly.
- [3] Spanhol, F., Cavalin, P., Oliveira, L. S., Petitjean, C., Heutte, L., Deep Features for Breast Cancer Histopathological Image Classification, 2017 IEEE International Conference on Systems, Man, and Cybernetics (IEEE SMC 2017), Banff, Canada, 2017
Some Statistics (updated Nov 30, 2022)
This dataset has been downloaded 7726 times from 141 different countries
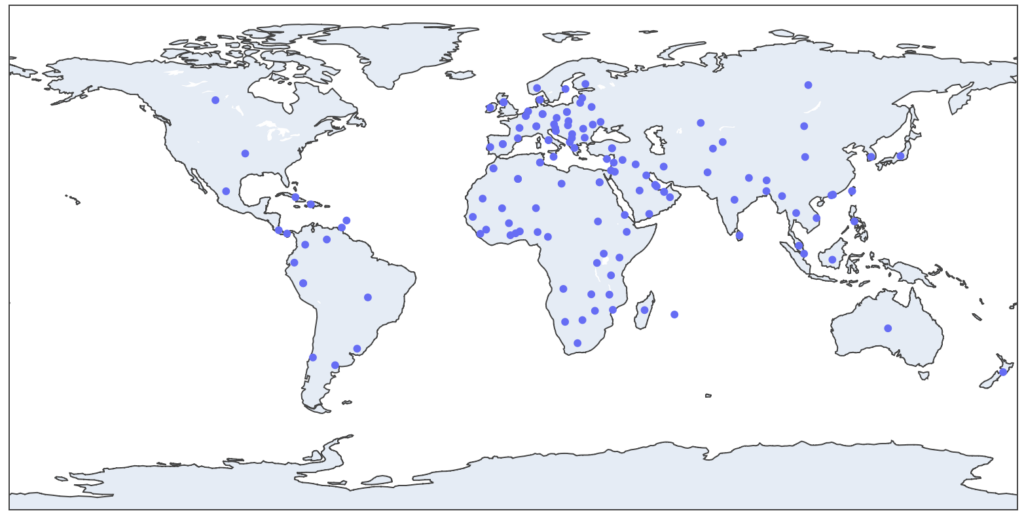
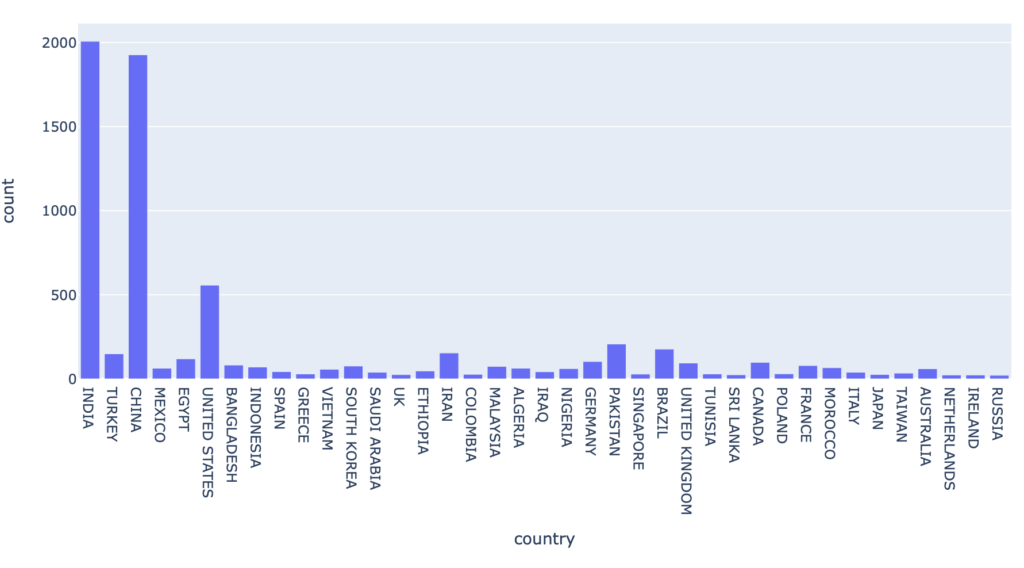
Countries with more than 30 downloads:
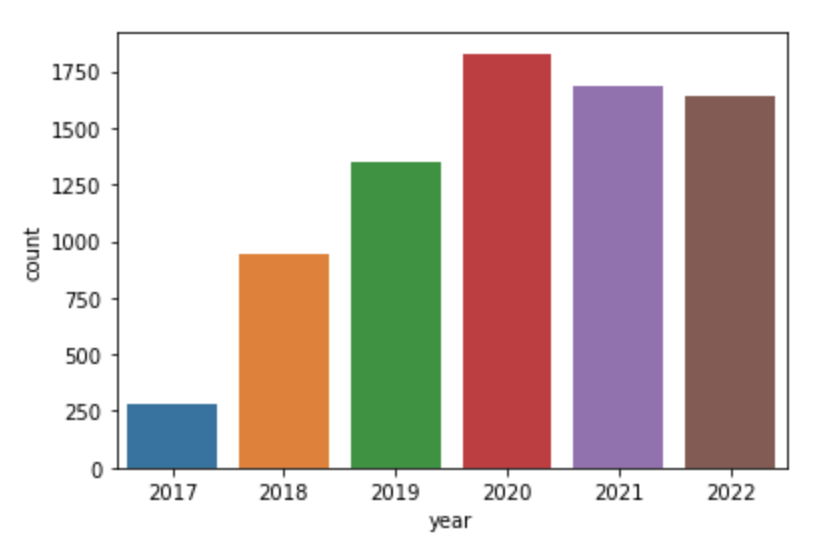
Downloads per year (since 2017)

BreaKHis – Breast Cancer Histopathological Database by Spanhol, F., Oliveira, L. S., Petitjean, C. and Heutte, L. is licensed under a Creative Commons Attribution 4.0 International License.
